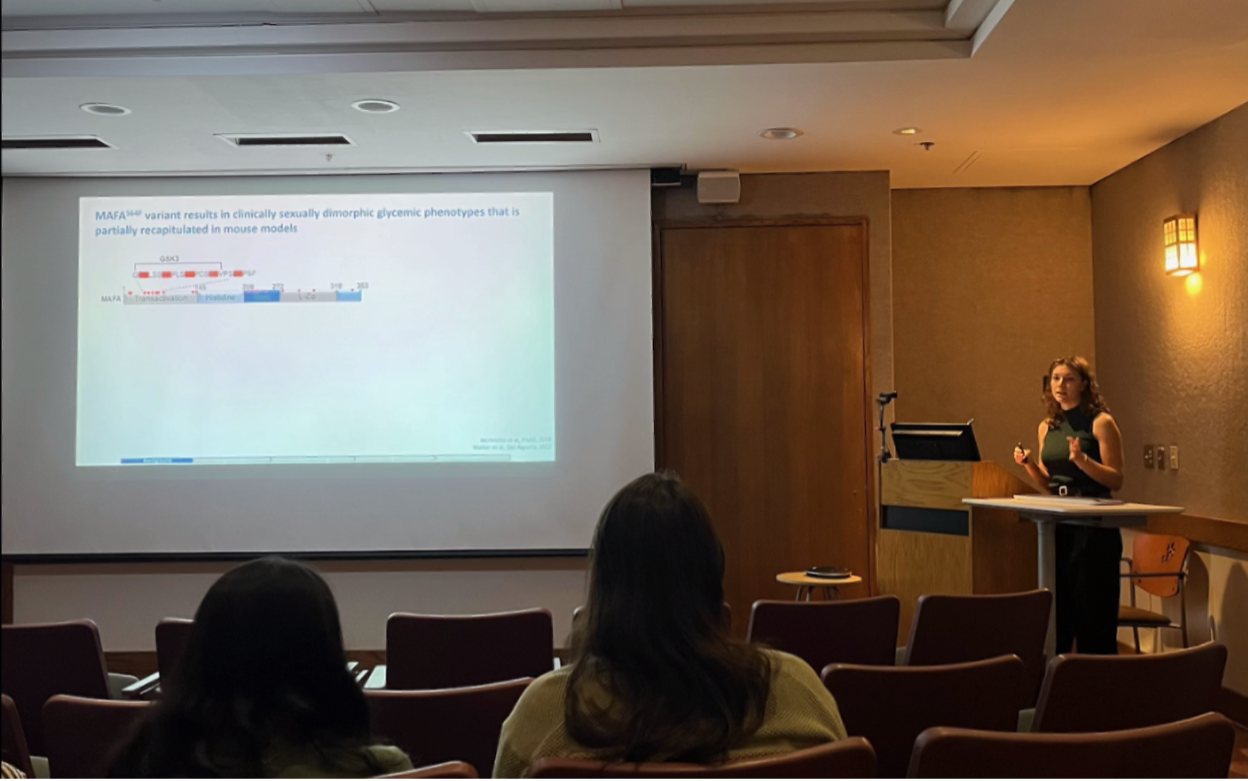
Effects of Mouse Genetic Background on the Phenotypic Presentation of MODY Variant MafAS64F
Manuscript in Preparation, 2025
Maurer, M., et al.
First-author manuscript characterizing genetic modifiers of diabetes using QTL mapping and multiomics approaches.
Discovery of 7-(Pyridin-3-yl)thieno[3,2-b]pyridine-5-carboxamides as NAMs of mGlu5
ACS Chemical Neuroscience [Submitted]
Henderson, S.H., Ringuette, A.E., Whomble, D.L., Captstick, R.A., Richardson, A.E., Maurer, M.A., et al.
This study describes the discovery of novel 7-(pyridin-3-yl)thieno[3,2-b]pyridine-5-carboxamides as mGlu5 negative allosteric modulators with enhanced pharmacological properties and selectivity profiles.
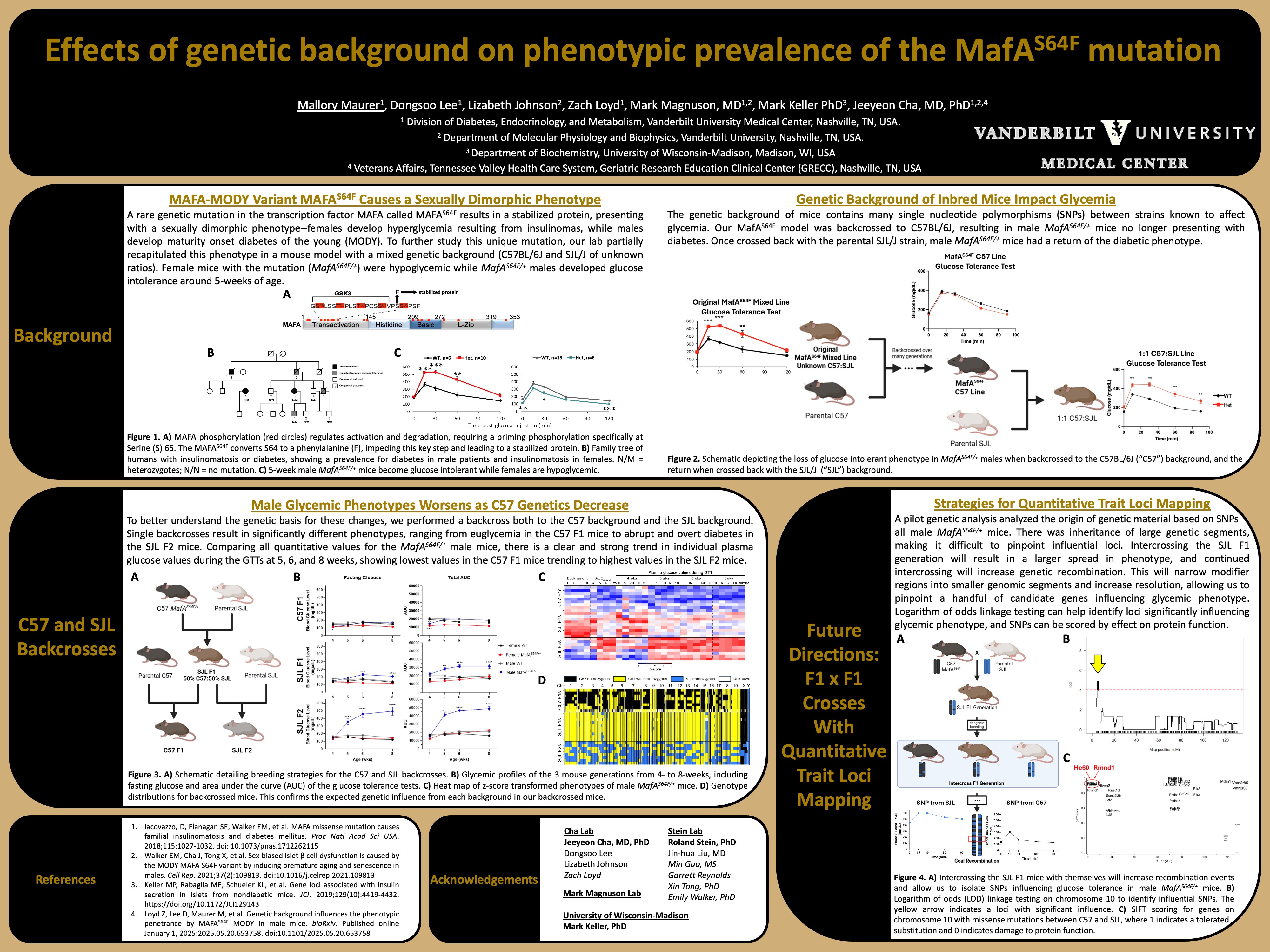
Genetic Background Influences the Phenotypic Penetrance by MAFA S64F MODY in Male Mice
bioRxiv [Preprint], 2025 Sep 15
Loyd, Z., Lee, D., Maurer, M., Buzzelli, L., Liu, J.H., et al.
This study demonstrates that genetic background significantly influences the phenotypic presentation of MAFA S64F MODY in male mice, revealing important genetic modifiers of diabetes pathogenesis.
Discovery of Thieno[3,2-b]pyridine-5-carboxamide and 2,3-Difluorobenzamide NAMs of mGlu5
ACS Med Chem Lett, 2025 Apr 22;16(5):865-874
Crocker, K.E., Henderson, S.H., Capstick, R.A., Whomble, D.L., Bender, A.M., Felts, A.S., Han, C., Engers, J.L., Billard, N.B., Maurer, M.A., et al.
This study identified thieno[3,2-b]pyridine-5-carboxamide and 2,3-difluorobenzamide as novel mGlu5 negative allosteric modulators that are highly potent, brain penetrant, and demonstrate improved oral bioavailability.
Discovery of 4-(5-Membered)Heteroarylether-6-methylpicolinamide NAMs of mGlu5
ACS Med Chem Lett, 2024 Nov 18;15(12):2210-2219
Childress, E.S., Capstick, R.A., Crocker, K.E., Ledyard, M.L., Bender, A.M., Maurer, M.A., et al.
This study developed novel mGlu5 negative allosteric modulators by replacing metabolic liabilities with 5-membered heterocycles, identifying VU6043653 as a highly brain penetrant and selective compound with improved pharmacological properties.
Development of VU6036864: A Triazolopyridine-Based High-Quality Antagonist Tool Compound of the M5 Muscarinic Acetylcholine Receptor
J Med Chem, 2024 Aug 22;67(16):14394-14413
Li, J., Orsi, D.L., Engers, J.L., Long, M.F., Capstick, R.A., Maurer, M.A., et al.
This study developed VU6036864, a potent and selective M5 muscarinic acetylcholine receptor antagonist with excellent brain penetration and oral bioavailability, advancing tool compounds for neurological research.
Evaluation of the Indazole Analogs of 5-MeO-DMT and Related Tryptamines as Serotonin Receptor 2 Agonists
ACS Med Chem Lett, 2024 Jan 19;15(2):302-309
Jayakodiarachchi, N., Maurer, M.A., Schultz, D.C., Dodd, C.J., Thompson Gray, A., et al.
This study synthesized and characterized novel indazole analogs of 5-MeO-DMT as potent serotonin receptor 2 agonists, examining their 5-HT2 pharmacology with emphasis on subtype selectivity.
Hyperglycemia Minimally Alters Primary Self-Renewing Human Colonic Epithelial Cells while TNFα Promotes Severe Intestinal Epithelial Dysfunction
Integr Biol (Camb), 2021 Jun 15;13(6):139-152
Dutton, J.S., Hinman, S.S., Kim, R., Attayek, P.J., Maurer, M., Sims, C.S., Allbritton, N.L.
This study investigated the effects of hyperglycemia and TNFα on primary human colonic epithelial cells, finding that TNFα was the dominant modulator of epithelial dysfunction compared to glucose levels.